 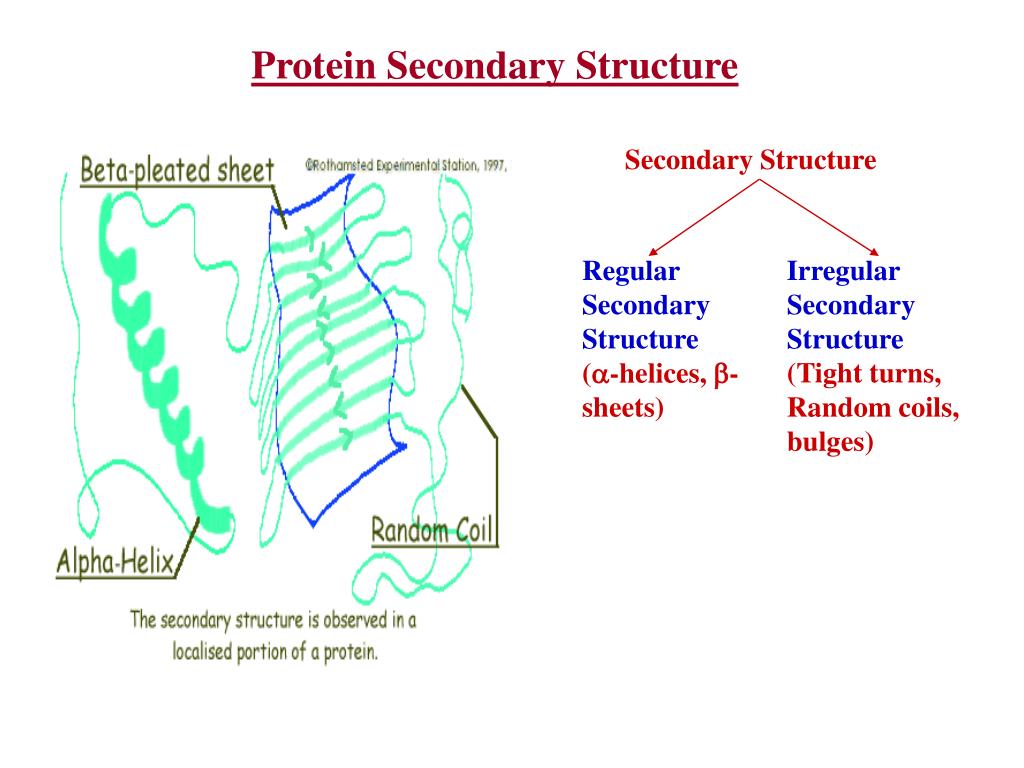
In that frame, tools for the geometric analysis of the protein graph, consisting of atoms and residues, are essential. Since three decades, the development of structural biology has been driven by the intention to relate the function of molecular objects to the physico-chemical rules at the atomic level. Due to its general modeling frame, Sequoia is able to handle graphs containing only C α atoms, which is particularly useful on low resolution structural input and in the frame of electron microscopy development. In contrast to the approaches classically used in structural biology, such as DSSP, Sequoia is able to capture the variations of geometry at the interface of adjacent secondary structure element.  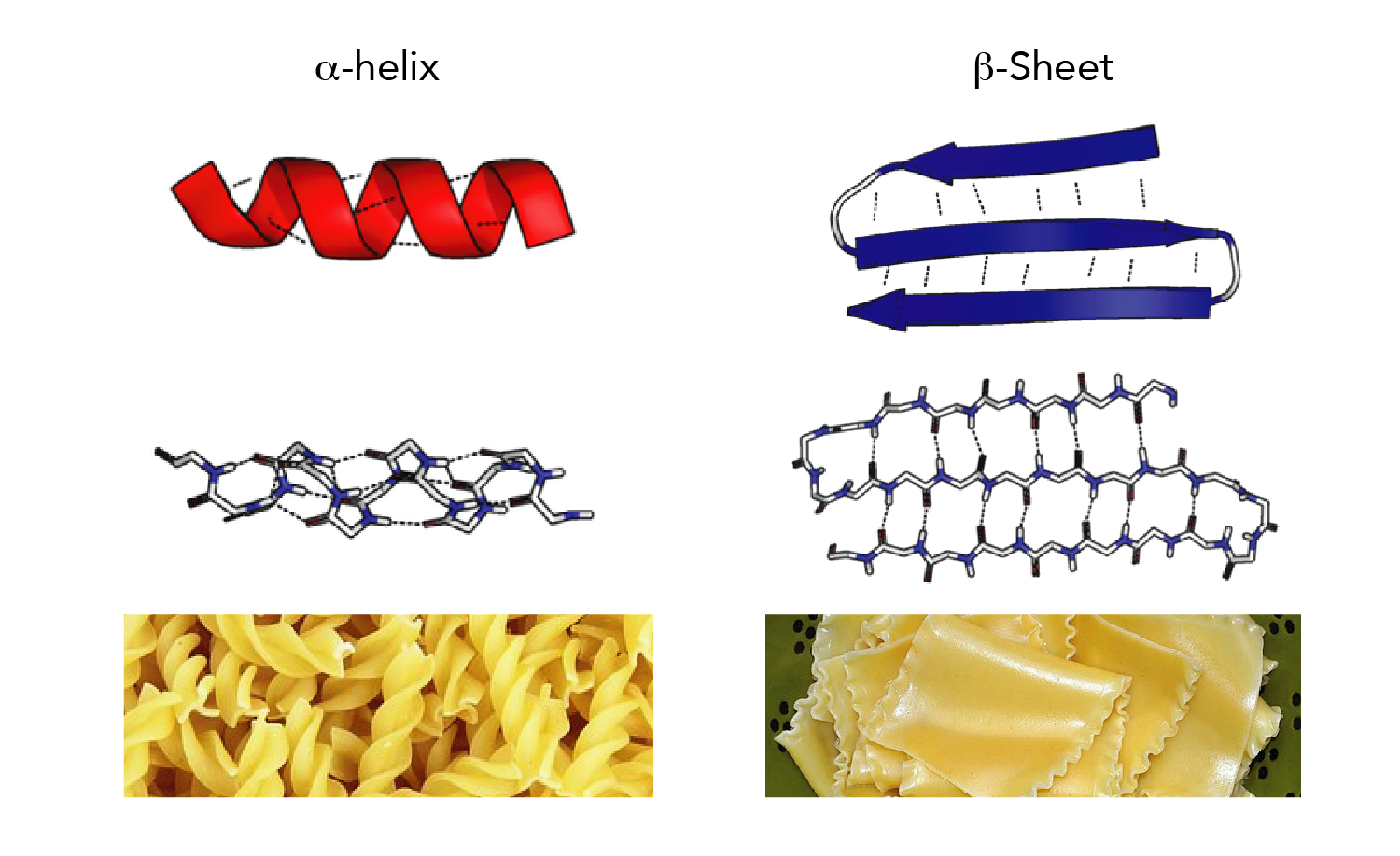
Sequoia permits the automatic detection of the secondary structure elements, with an F1-score larger than 80% for most of the cases, when α helices and β strands are predicted. Any pair of residues, independently of its covalent bonds along the primary sequence of the protein, is tagged with this distance and angle information. This neural network takes as input the topology of the given protein graph, where the vertices are protein residues, and the edges are weighted by values of distances and pseudo-dihedral angles generalizing the backbone angles ϕ and ψ. 
Our method uses a message passing neural network, named Sequoia, which allows the automatic prediction of secondary structure elements from the values of local distances and angles between the protein atoms. We introduce a new methodology to estimate the secondary structure elements from the values of local distances and angles between the protein atoms. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
